In the past, I have done quite a lot of things related to text classification and I did some writing about it. Now came a project where I needed to build a named entity recognizer. Since I had a couple of hour (almost a working day) struggle to understand how it all goes together, here is a tutorial (for both my reference and someone’s learning).
What is NER?
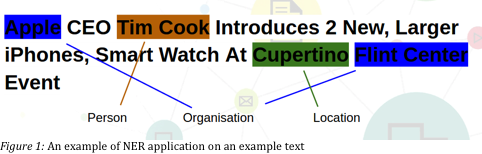
Named entity recognizer is a program that recognizes named entity in text. The named entities could be anything from locations, company or person’s names, drug or disease names, etc. Bellow is an image of named entities recognized in a sentence about Tim Cook.

If we go to the basics, it is a text classification, however, in this case we classify each word whether it is some of the named entities of interest or is it not. However, we are looking at the sequence of words and sentence or even whole text as a sequence. The words that occurred in the sequence will influence our classification, and therefore we need to use sequence modelling algorithms. State of the art at the moment is with word embeddings and deep neural network, however, here we will review how to build a NER using simpler algorithms such as conditional random fields (CRF).
CRF-based NER for de-identification
For the purpose of this tutorial, we will use de-identification dataset that was released for I2B2 clinical challenge. You can request the access to the dataset here: https://www.i2b2.org/NLP/DataSets/
I2B2 files are XML files, containing text and at the bottom a set of annotations, where it states the offset and classes in XML tags. The example is:
<DATE id="P0" start="16" end="26" text="2067-05-03" TYPE="DATE" comment="" />
<AGE id="P1" start="50" end="52" text="55" TYPE="AGE" comment="" />
<NAME id="P2" start="290" end="296" text="Oakley" TYPE="DOCTOR" comment="" />
We will be trying to predict the TYPE part of this XML.
Firstly, we need to read the files. We do it using the following function:
def readSurrogate(path):
onlyfiles = [f for f in listdir(path) if isfile(join(path, f))]
documents = []
for file in onlyfiles:
tree = ET.parse(path+"/"+file)
root = tree.getroot()
document_tags = []
for child in root:
if child.tag == "TEXT":
text = child.text
if child.tag == "TAGS":
for chch in child:
tag = chch.tag
attributes = chch.attrib
start = attributes["start"]
end = attributes["end"]
content = attributes["text"]
type= attributes["TYPE"] document_tags.append({"tag":tag,"start":start,"end":end,"text":content,"type":type})
documents.append({"id":file,"text":text,"tags":document_tags})
return documents
The presented function makes document structure that contains text and a list of tags as the document. However, this is not overly useful for CRF, as it takes a sequences of words with labels for training. In testing phase, it as well takes sequences of words and predicts labels (e.g. age, name, etc).
The sequence in this sense should not be long and usually is taken to represent a sentence, while sometimes whole documents would be used. However, you should note that longer sequence will require more memory usage and more processing. Here I did my quite a big error that costed quite some number of hours of work. I have concatenated whole of all documents in the dataset (about 5MB of XML files, about 600 files) to a single sequence and tried to train with that. That did not work, as on Windows it was throwing out of memory exception, while on Linux (where I did primarily this code), it was not throwing exception, but utilized all memory to the point that computer was unusable and needed a restart.
Now, in order to create sequences, I have ended up using the following code:
def tokenize_fa(documents):
sequences = []
sequence = []
for doc in documents:
if len(sequence)>0:
sequences.append(sequence)
sequence = []
text = doc["text"]
file = doc["id"]
text = text.replace("\"", "'")
tokens = custom_span_tokenize(text)
for token in tokens:
token_txt = text[token[0]:token[1]]
found = False
for tag in doc["tags"]:
if int(tag["start"])<=token[0] and int(tag["end"])>=token[1]:
token_tag = tag["tag"]
found = True
if found==False:
token_tag = "O"
sequence.append((token_txt,token_tag))
if token_txt == ".":
sequences.append(sequence)
sequence = []
sequences.append(sequence)
return sequences
def custom_word_tokenize(text, language='english', preserve_line=False):
tokens = []
sentences = [text] if preserve_line else nltk.sent_tokenize(text, language)
for sent in sentences:
for token in _treebank_word_tokenizer.tokenize(sent):
if "-" in token:
m = re.compile("(\d+)(-)([a-zA-z-]+)")
g = m.match(token)
if g:
for group in g.groups():
tokens.append(group)
else:
tokens.append(token)
else:
tokens.append(token)
return tokens
def custom_span_tokenize(text, language='english', preserve_line=False):
tokens = custom_word_tokenize(text)
return align_tokens(tokens, text)
There are a couple of things to unpack. The function called custom_span_tokenize takes tokens that are created using custom_word_tokenize and makes spans. On the other hand, custom_word_tokenize is a custom tokenizer that is able to split in a right way phrases, such as 47-year-old man, as in this dataset, only 47 is annotated as age. The function tokenize_fa makes a sequences of tokens with their label as tuple like (‘right’,’O’).
Now we can split the dataset and create sequences out of it. We can do it in the following way:
train_docs = documents[:400]
test_docs = documents[400:]
print("Tokenizing")
train_sequences = tokenize_fa(train_docs)
test_sequences = tokenize_fa(test_docs)
print("Tokenized")
The next bit is creation of datasets for training and testing. Training dataset should have 2 components: a sequence of tokens with other features about them (X) and a sequence of labels (y). For testing we do the same, so we can later compare real y and predicted y.
crf = CRF_DeId_NER()
crf.X_train = []
crf.y_train = []
crf.X_test = []
crf.y_test = []
print("Training set creation")
for seq in train_sequences:
features_seq = []
labels_seq = []
for i in range(0,len(seq)):
features_seq.append(crf.word2features(seq, i))
labels_seq.append(crf.word2labels(seq[i]))
crf.X_train.append(features_seq)
crf.y_train.append(labels_seq)
print("Training set created")
print("Testing set creation")
for seq in test_sequences:
features_seq = []
labels_seq = []
for i in range(0,len(seq)):
features_seq.append(crf.word2features(seq, i))
labels_seq.append(crf.word2labels(seq[i]))
crf.X_test.append(features_seq)
crf.y_test.append(labels_seq)
print("Testing set created")
In this code, we iterate through both training and testing sequence and generate feature set sequence and label sequence. This is done in our case inside my CRF_DeId_NER class. Building labels is pretty simple:
def word2labels(self, sent):
return sent[1]
However, creating features for CRF is the complex process and there is a whole field of feature engineering around this. Here is what we used:
def shape(self,word):
shape = ""
for letter in word:
if letter.isdigit():
shape = shape + "d"
elif letter.isalpha():
if letter.isupper():
shape = shape + "W"
else:
shape = shape + "w"
else:
shape = shape + letter
return shape
def word2features(self,sent, i):
word = sent[i][0]
#postag = sent[i][1]
features = {
'bias': 1.0,
'word.lower()': word.lower(),
'word.isupper()': word.isupper(),
'word.istitle()': word.istitle(),
'word.isdigit()': word.isdigit(),
'word.shape()':self.shape(word),
'word.isalnum()':word.isalnum(),
'word.isalpha()':word.isalpha(),
}
if i > 0:
word1 = sent[i - 1][0]
#postag1 = sent[i - 1][1]
features.update({
'-1:word.lower()': word1.lower(),
'-1:word.istitle()': word1.istitle(),
'-1:word.isupper()': word1.isupper(),
'-1:word.isdigit()': word1.isdigit(),
'-1:word.isalnum()':word1.isalnum(),
'-1:word.isalpha()':word1.isalpha(),
})
else:
features['BOS'] = True
if i > 1:
word2 = sent[i - 2][0]
features.update({
'-2:word.lower()': word2.lower(),
'-2:word.istitle()': word2.istitle(),
'-2:word.isupper()': word2.isupper(),
'-2:word.isdigit()': word2.isdigit(),
'-2:word.isalnum()': word2.isalnum(),
'-2:word.isalpha()': word2.isalpha(),
# '-2:postag': postag2,
# '-2:postag[:2]': postag2[:2],
})
else:
features['BOS1'] = True
if i > 2:
word3 = sent[i - 3][0]
#postag3 = sent[i - 3][1]
features.update({
'-3:word.lower()': word3.lower(),
'-3:word.istitle()': word3.istitle(),
'-3:word.isupper()': word3.isupper(),
'-3:word.isdigit()': word3.isdigit(),
'-3:word.isalnum()': word3.isalnum(),
'-3:word.isalpha()': word3.isalpha(),
})
else:
features['BOS2'] = True
if i > 3:
word4 = sent[i - 4][0]
features.update({
'-4:word.lower()': word4.lower(),
'-4:word.istitle()': word4.istitle(),
'-4:word.isupper()': word4.isupper(),
'-4:word.isdigit()': word4.isdigit(),
'-4:word.isalnum()': word4.isalnum(),
'-4:word.isalpha()': word4.isalpha(),
})
else:
features['BOS2'] = True
if i < len(sent) - 1:
word1 = sent[i + 1][0]
features.update({
'+1:word.lower()': word1.lower(),
'+1:word.istitle()': word1.istitle(),
'+1:word.isupper()': word1.isupper(),
'+1:word.isdigit()': word1.isdigit(),
'+1:word.isalnum()': word1.isalnum(),
'+1:word.isalpha()': word1.isalpha(),
})
else:
features['EOS'] = True
if i < len(sent) - 2:
word12 = sent[i + 2][0]
features.update({
'+2:word.lower()': word12.lower(),
'+2:word.istitle()': word12.istitle(),
'+2:word.isupper()': word12.isupper(),
'+2:word.isdigit()': word12.isdigit(),
'+2:word.isalnum()': word12.isalnum(),
'+2:word.isalpha()': word12.isalpha(),
})
else:
features['EOS2'] = True
if i < len(sent) - 3:
word13 = sent[i + 3][0]
#postag13 = sent[i + 3][1]
features.update({
'+3:word.lower()': word13.lower(),
'+3:word.istitle()': word13.istitle(),
'+3:word.isupper()': word13.isupper(),
'+3:word.isdigit()': word13.isdigit(),
'+3:word.isalnum()': word13.isalnum(),
'+3:word.isalpha()': word13.isalpha(),
})
else:
features['EOS2'] = True
if i < len(sent) - 4:
word14 = sent[i + 4][0]
features.update({
'+4:word.lower()': word14.lower(),
'+4:word.istitle()': word14.istitle(),
'+4:word.isupper()': word14.isupper(),
'+4:word.isdigit()': word14.isdigit(),
'+4:word.isalnum()': word14.isalnum(),
'+4:word.isalpha()': word14.isalpha(),
})
else:
features['EOS2'] = True
return features
For each word in a sequence that we look at, we figure out whether that word has first capital letter, what is the word lower-cased, whether all letters are upper cased, whether it is digit, whether it is alphanumberic, or just alpha. We look for all these characteristics in surrounding 4 words on both sides. For the word we look at, we also evaluate shape of it (whether it has some shape pattern).
Now when we extracted features, we can train the algorithm. Training is done using the following function:
def train(self):
self.crf_model = sklearn_crfsuite.CRF(
algorithm='lbfgs',
c1=0.1,
c2=0.05,
max_iterations=200,
all_possible_transitions=True
)
self.crf_model.fit(self.X_train, self.y_train)
print("Training")
crf.train()
print("Train end")
At the end model can be evaluated, using the following code:
labels = list(crf.crf_model.classes_)
labels.remove('O')
print(labels)
y_pred = crf.crf_model.predict(crf.X_test)
f1_score = metrics.flat_f1_score(crf.y_test, y_pred,
average='weighted', labels=labels)
precision_score = metrics.flat_precision_score(crf.y_test, y_pred,
average='weighted', labels=labels)
recall_score = metrics.flat_recall_score(crf.y_test, y_pred,
average='weighted', labels=labels)
stats = metrics.flat_classification_report(crf.y_test, y_pred,
labels=labels)
print("Precision: "+str(precision_score))
print("Recall: "+str(recall_score))
print("F1-score: "+str(recall_score))
print(stats)
filename = '../Models/crf_baseline_model.sav'
pickle.dump(crf.crf_model, open(filename, 'wb'))
print("Done with all")
The score that can be obtained with the algorithm is the following:
precision recall f1-score support
DATE 0.97 0.93 0.95 1124
AGE 0.96 0.85 0.90 159
NAME 0.96 0.83 0.89 1274
LOCATION 0.91 0.63 0.74 599
PROFESSION 0.60 0.09 0.15 69
ID 0.96 0.71 0.81 112
CONTACT 0.93 0.88 0.90 89
PHI 0.00 0.00 0.00 0
avg / total 0.95 0.81 0.87 3426
The full code can be seen at:
https://github.com/nikolamilosevic86/NERo/blob/master/DeID/DeID_CRF_based.py
Or under NERo project:
https://github.com/nikolamilosevic86/NERo